Most Recent in ArcGIS Blog
Updated U.S. geographic boundaries in ArcGIS Business Analyst (June 2024)
With the June 2024 data release, there are boundary changes to a few U.S. geography types used in ArcGIS Business Analyst Web App.
Identify the best location for an urgent care center
Multiple Authors | Analytics | April 2, 2024
Use suitability analysis in ArcGIS Business Analyst Web App to locate a site for a new urgent care center in Maverick County, Texas.
Most Recent in ArcGIS Blog
Multiple Authors | ArcGIS Business Analyst | May 10, 2024
With the June 2024 data release, there are boundary changes to a few U.S. geography types used in ArcGIS Business Analyst Web App.

Multiple Authors | ArcGIS Field Maps | May 10, 2024
Q&A from the Maximize field efficiecy with Field Maps webinar.
Chris Wesson | ArcGIS Pro | May 9, 2024
Cartographic finishing for polygon marker symbol layers with the new Convert marker placement to points tool.

Stephen Kredel | ArcGIS AllSource | May 9, 2024
ArcGIS AllSource 1.2 has released and with it comes Simplified Feature Creation, Military Overlay, and ArcGIS Video Server connectivity.
Hong Xu | ArcGIS Pro | May 9, 2024
Learn how to explore, manage, and analyze EMIT hyperspectral imagery using ArcGIS Pro 3.3

Tanner Yould | Developers | May 9, 2024
ArcGIS Maps SDK for Native Apps version 200.4 adds support for 3D Tiles layers!
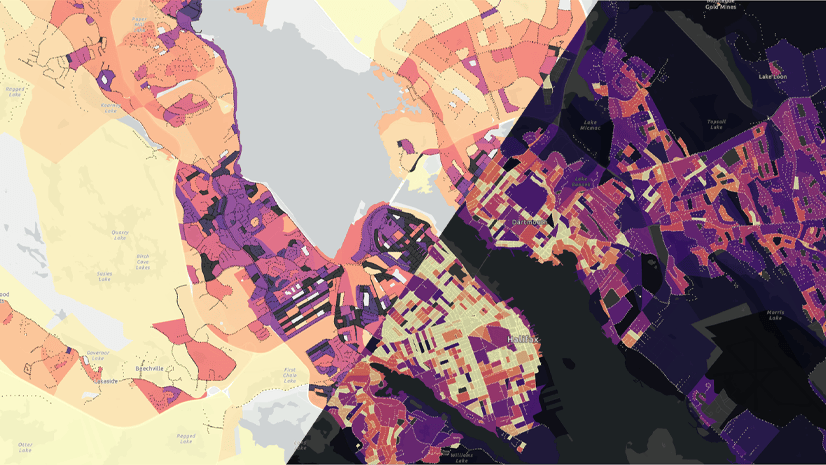

Heather Smith | ArcGIS Online | May 9, 2024
Watch this short video to learn how to choose color schemes that work well with light or dark basemaps.

Multiple Authors | ArcGIS Pro | May 9, 2024
What's new for GeoAI in the Image Analyst extension of ArcGIS Pro 3.3

Matvei Stefarov | ArcGIS Maps SDK for .NET | May 8, 2024
Release 200.4 adds DateOnly, TimeOnly, and TimestampOffset fields to provide flexible temporal data storage and enhance time zone support.
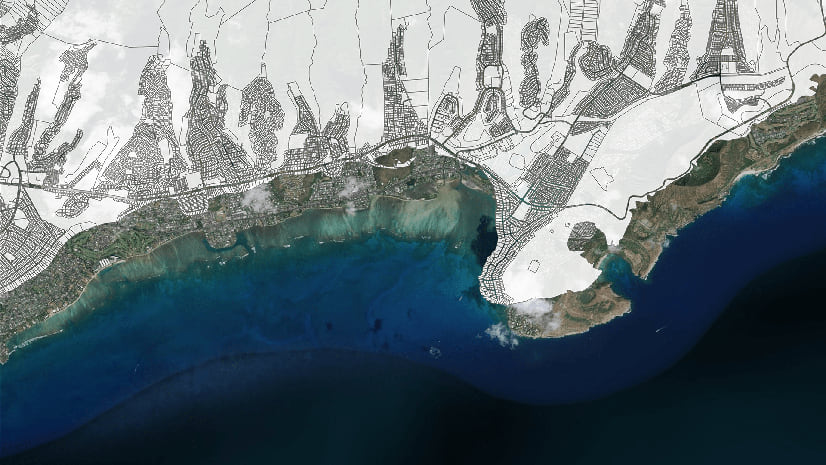

Bojan Šavrič | ArcGIS Pro | May 8, 2024
The new U.S. datums of 2022 will soon be released. This article covers what is coming and how you should prepare your data.
Lauryn Carey | ArcGIS Business Analyst | May 8, 2024
Read a summary of the newest features, enhancements, and improvements in the ArcGIS Business Analyst Pro May Release.
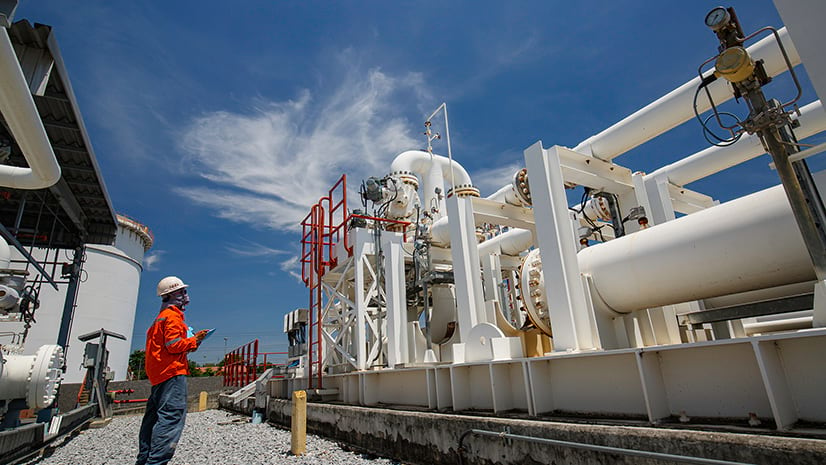

Mark Barker | ArcGIS Drone2Map | May 8, 2024
Learn how inspection workflows are used in ArcGIS Drone2Map to monitor assets and assist in decision making.
Krista Carlson | ArcGIS Pro | May 7, 2024
ArcGIS Bathymetry introduces three new tools and enhances Compose Surface capabilities in ArcGIS Pro 3.3

Multiple Authors | ArcGIS Utility Network | May 7, 2024
Learn about the new capability of the utility network to use the digitized direction to perform analysis of stormwater and wastewater networks!

Multiple Authors | ArcGIS Pro | May 7, 2024
Model realistic water flow, share your GIS story, add hyperlinks to your text elements, and so much more with the ArcGIS Pro 3.3 release.
Multiple Authors | ArcGIS Hub | May 7, 2024
Laura Sharp, PhD, shares how to capture residents’ perspectives and unique needs into the City of Tucson's decision-making processes.

William Hackney | ArcGIS StoryMaps | May 7, 2024
Learn about the options available for configuring your web maps and scenes in ArcGIS StoryMaps to make them effective and impactful.

Dejan Parac | ArcGIS IPS | May 7, 2024
Learn what's new in the ArcGIS IPS May 2024 release, explore native GIS support, IPS aware maps and ArcGIS IPS Setup app enhancements.

Multiple Authors | ArcGIS Hub | May 7, 2024
Building local resilience against community challenges means collaborating with your community early.

Multiple Authors | ArcGIS Utility Network | May 6, 2024
2024 Network management release plan for Utilities and Telecom industries

Thomas Coughlin | ArcGIS Experience Builder | May 6, 2024
Group filters are new with the February 2024 update. Available now.

Jon DeRose | ArcGIS Utility Network | May 6, 2024
Learn more about exciting new functionality and improvements made with the recent Network Management Release of the ArcGIS Utility Network

Multiple Authors | ArcGIS Enterprise | May 6, 2024
Discussion with Xander Bakker, Esri Colombia, on his unconventional path to GIS and how he truly loves his work.

Robert Krisher | ArcGIS Utility Network | May 5, 2024
Learn how to model flow and perform analysis with wastewater networks using the ArcGIS Utility Network

Caitlyn North | ArcGIS QuickCapture | May 3, 2024
The answers to questions from the March 2024 ArcGIS QuickCapture Introduction and What's New Webinar.

Multiple Authors | ArcGIS Video Server | May 2, 2024
The 11.3 release of ArcGIS Video Server has various improvements that expand how customers can interact with video in ArcGIS.
